Pralsetinib Polypharmacology & Toxicity Knowledge Graph
A Neuro-Symbolic Approach to Drug Safety using Knowledge Graphs.
Predicting Unknown Side Effects of the Cancer Drug Pralsetinib
A Neuro-Symbolic Approach to Drug Safety using Knowledge Graphs
Team Members: Ryan Cao, Suchit Bhayani, Ishaan Bal, Taranvir Chima
Mentors: Raju Pusapati (Solix), Murali Krishnam (Solix), Justin Eldridge (UCSD)
Abstract
Pralsetinib has limited long-term real-world safety data. Post-marketing pharmacovigilance already shows unexpected adverse effects beyond on-target effects. Because it is new, used in a small subset of patients, and has complex kinase biology, there is a high “unknown off-target space.” Our approach builds a knowledge graph linking mechanisms to toxicity, enriching it with biological pathways, and trains an AI model to predict specific off-target adverse outcomes.
Introduction: The Problem
- The Hook: Modern pharma R&D is constrained by fragmented, heterogeneous evidence. Assays, literature, clinical reports, and observational data rarely align in a single structured representation.
- The Problem: Pralsetinib, a recently approved cancer drug, needs better mapping of its off-target effects to predict unexpected adverse side effects like severe infections and cognitive disorders.
- Our Goal: We shift from text-only extraction to a graph-based representation. By building a Knowledge Graph (KG) that encodes explicit relationships, we enable auditable reasoning and reproducible off-target hypotheses.
Objectives
- Map Evidence: Consolidate fragmented biomedical data (assays, clinical trials, literature) into a unified Knowledge Graph.
- Enrich with Biology: Use established ontologies (Gene Ontology, CTD) to link proteins to broader biological pathways and clinical outcomes.
- Predict Unknowns: Train an AI model to discover missing links, predicting novel side effects based on multi-hop graph structures rather than direct connections alone.
Methods
1. Knowledge Graph Construction
We constructed the baseline KG from PubChem-derived sources:
- Bioactivity assays (direct drug-target evidence)
- Clinical trials and indications (disease treatments)
- Literature mining (adverse event text mining)
- Chemical/gene co-occurrence signals (weak evidence links)
Visualizing the Graph Structure Initially, our graph was highly centralized around Pralsetinib, relying heavily on chemical and gene co-occurrences. After enriching it with biological ontologies, the composition shifted dramatically.
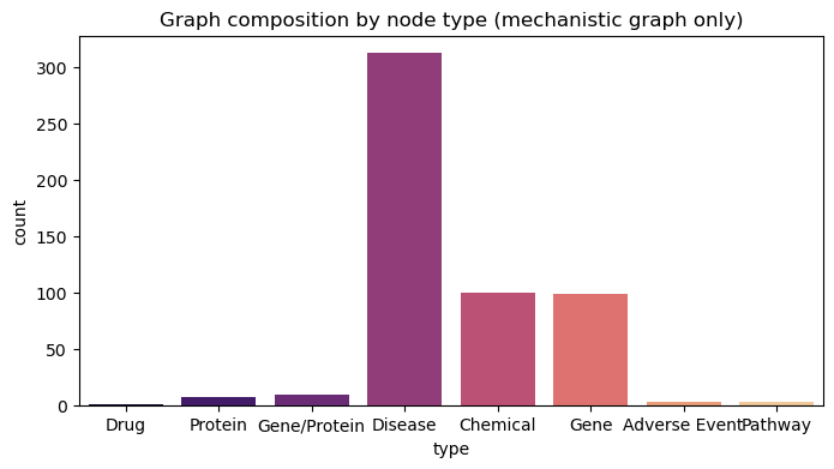 Above: Node type distribution showing how proteins and diseases become the dominant entities after our ontology enrichment, significantly increasing the graph’s mechanistic breadth.
Above: Node type distribution showing how proteins and diseases become the dominant entities after our ontology enrichment, significantly increasing the graph’s mechanistic breadth.
Visualizing this network highlights the complexity of drug interactions, but also reveals a structural challenge:
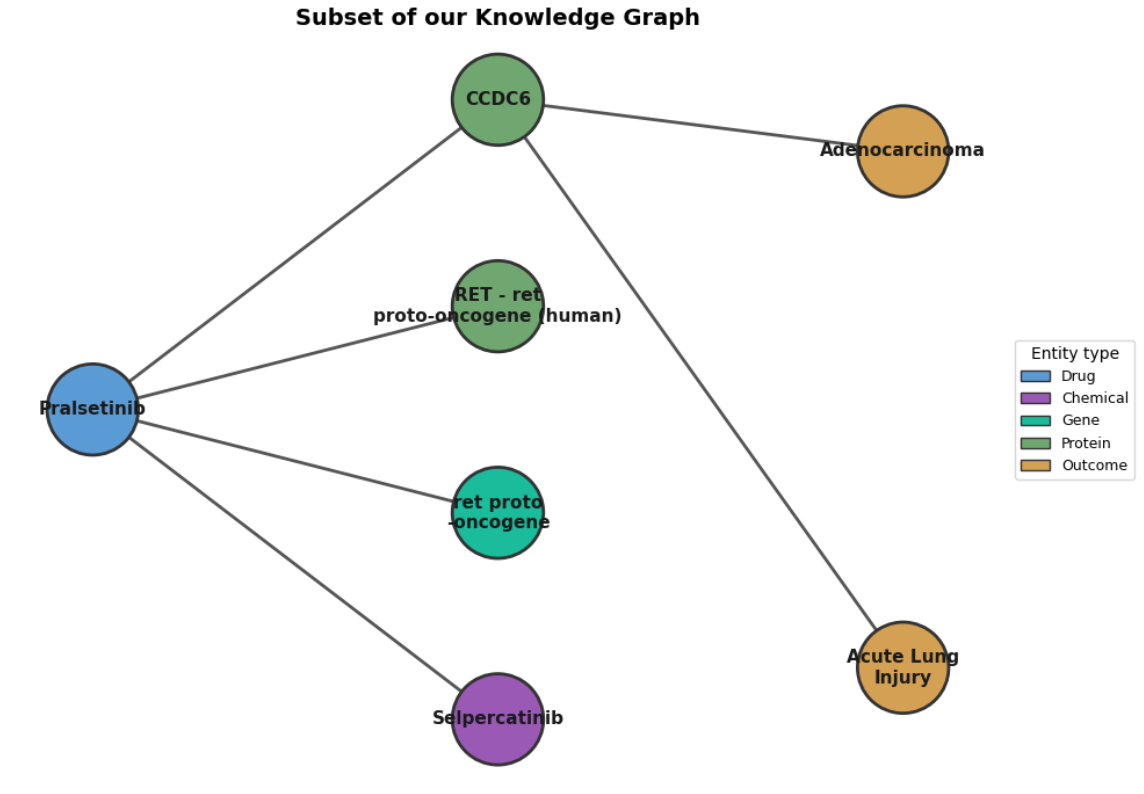 Above: A subset visualization of the Knowledge Graph. Notice how heavily centralized the edges remain around Pralsetinib (forming a “star topology”). This centralization motivates our use of Graph Neural Networks to look beyond direct edges and find hidden lateral connections.
Above: A subset visualization of the Knowledge Graph. Notice how heavily centralized the edges remain around Pralsetinib (forming a “star topology”). This centralization motivates our use of Graph Neural Networks to look beyond direct edges and find hidden lateral connections.
Click here for the technical enrichment details
To move beyond a simple "star topology" centered on the drug, we enriched the KG using Biomedical Ontologies:
- Gene Ontology (GO): We queried GO via the UniProt API to link proteins to Biological Processes, adding
involved_inedges and Pathway nodes. - Comparative Toxicogenomics Database (CTD): We used CTD to connect proteins to clinical outcomes (Disease/AE) via
associated_withedges, focusing on oncology to prevent node explosion.
After enrichment, the graph grew to 580 nodes and 1,456 edges, with proteins and diseases dominating the structure.
2. Modeling
We compared a simple baseline model against an advanced AI model (Graph Neural Network).
- Baseline: Ranks proteins using only direct drug-protein connections.
- AI Model: Looks at the “big picture” (the full KG), learning from shared connectivity patterns and multi-hop structures to make predictions.
Click here for more model details
Our GNN uses the full KG, learns node embeddings with a 2-layer Graph Convolutional Network (GCN), and scores links with a Multi-Layer Perceptron (MLP) head. It is trained to predict (Pralsetinib, inhibits, Protein) and optionally (Protein, associated_with, Outcome).
Results
To evaluate our approach, we compared how well our AI model (GNN) and the baseline model could “rediscover” known targets that we intentionally hid during training.
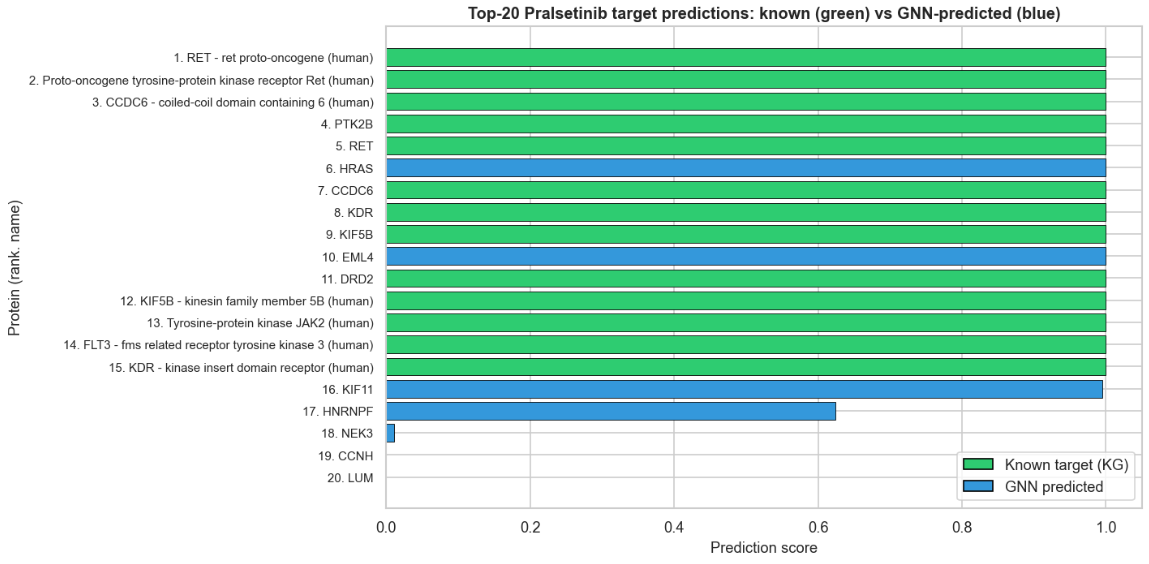 Above: Known targets recovered in the top-k predictions. The GNN exploits multi-hop structures to find hidden connections much more effectively than the baseline.
Above: Known targets recovered in the top-k predictions. The GNN exploits multi-hop structures to find hidden connections much more effectively than the baseline.
- High-Level Explanation: When evaluated on known targets, our AI model successfully recovered 100% of the known drug-protein targets within its top 20 predictions!
- Beating the Baseline: The AI retrieved significantly more known targets at larger k (Top-5, Top-10, Top-20) compared to the baseline model, highlighting the value of multi-hop graph structure over direct evidence alone.
Conclusion & Next Steps
Conclusion: Our pipeline demonstrates that combining structured assay evidence with biological pathways enables mechanistic reasoning. The final deliverable provides pharmacologists with transparent, data-backed hypotheses for Pralsetinib’s safety-relevant outcomes, addressing the critical interpretability challenges in AI-driven drug discovery.
Next Steps / Future Directions:
- Expand the Protein Set: Performance is currently constrained by the limited number of known positives (13 total). Expanding this set will allow for more robust future evaluations.
- Hold-out Testing: Conduct larger held-out evaluations for both baseline and AI models to ensure fair comparisons.
Get in Touch
GitHub: Ishaanbal/DSC180B-B23-Knowledge-Graph-and-Biomedical-Ontology
- Contact:
- Ryan Cao – rycao@ucsd.edu
- Suchit Bhayani – sbhayani@ucsd.edu
- Ishaan Bal – ibal@ucsd.edu
- Taranvir Chima – tchima@ucsd.edu
We welcome any questions or suggestions for further exploration.